Back in 2020 we found potential signs of life on Venus. Using the James Clerk Maxwell Telescope in Hawaii and the Atacama Large Millimeter/submillimeter Array in Chile a team of researchers used rotational absorption spectroscopy to investigate the Venusian atmosphere. They were stunned to find a phosphine signal. Stunned because there was no known abiotic process which produces phosphine outside of industrial processes: it’s something only produced by life as far as we know. The discovery either meant there was some unknown geological process which can produce phosphine – certainly of interest to a few people – or that there was some kind of life in the atmosphere. Stunning indeed.
However, the team were cautious and very open to the possibility they had missed something. But it was less what they missed, more what they added.

Image Credit: William Montgomerie, EAO/JCMT. There’s some cool images of it here.
Creating phosphine from noise
They explored a range of polynomials ranging from 14 to 1, finding that 8 worked best for the JCTM data and 12 for the ALMA data. But there’s a whole world in the word ‘best’. Reading the original paper I found it quite difficult to decipher what best meant so I’ll quote directly:
“The flattest spectral baselines were obtained by restricting the passband to 40 km/s from Venus’ velocity, and interpolating across |v| = 5 km/s. Polynomials of 12th -order were necessary, higher than the 8th – order JCMT fits; we verified that lower orders, e.g. 11, yielded worse noise while higher orders, e.g. 14, did not improve the noise.”
I take that to mean they visually inspected the spectra themselves, and felt the respective polynomials they subtracted made for the flattest baseline. This is far from my field, but inspecting the original and corrected spectra raises an eyebrow. Notice below (left panel) how the polynomial (dotted blue line) diverges from the raw data (solid blues lines) around 0. The right panel the shows the ‘corrected’ spectrum with quite the dip. It seems clear that the estimated polynomial baseline is not tracking the baseline.
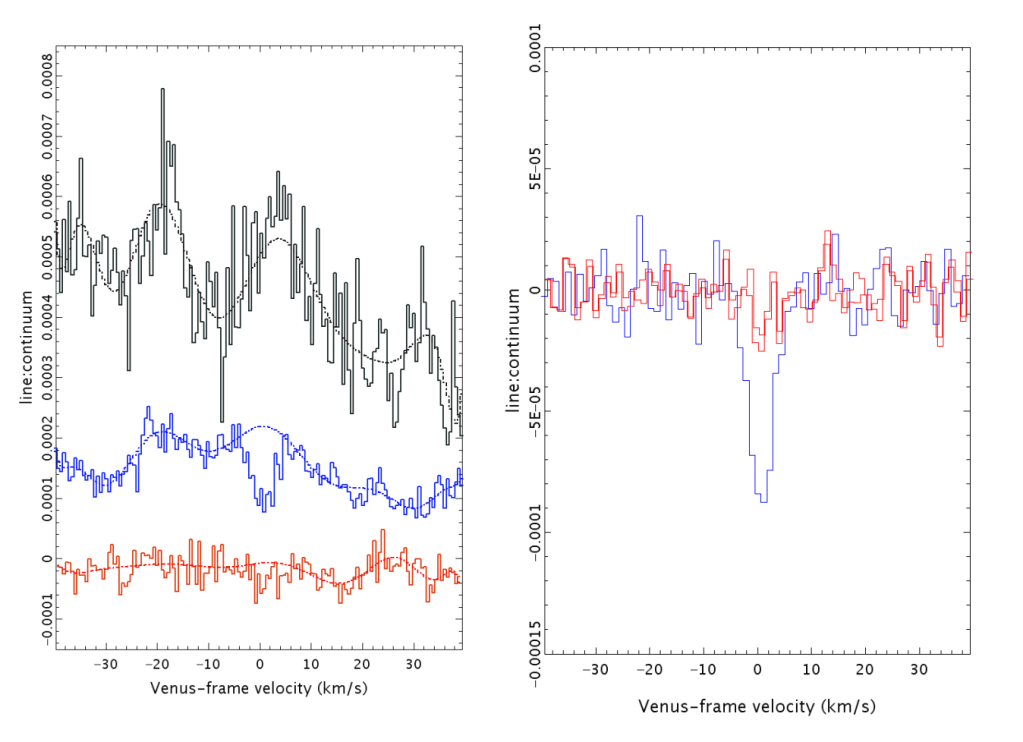
Supplementary figure 4. This is from the ALMA data. Something similar is observed in the JCMT data.
A reanalysis of the data by another team found that indeed the choice of polynomial invented a signal from noise. Interestingly, they found a number of other polynomial choices could reveal spurious signals.
This whole story is actually one of good science. True the reviewers didn’t flag it, but the devil was found in the details post publication. And I feel there’s a lesson to be had for the vibrational spectroscopy community.
Vibrational Spectroscopy
I came across this story during my PhD. in Raman spectroscopy. Everyone baseline corrected their data. Everyone. You have to, else how do you account for the fluorescence signal in your data? It’s just what you have to do with biomedical spectra. There’s a vast array of methods for baseline correction, and each with many hyperparameters. And none with any principled way of choosing between them. You would just eyeball the data until it looked ‘good’. Below shows just a fraction of methods.
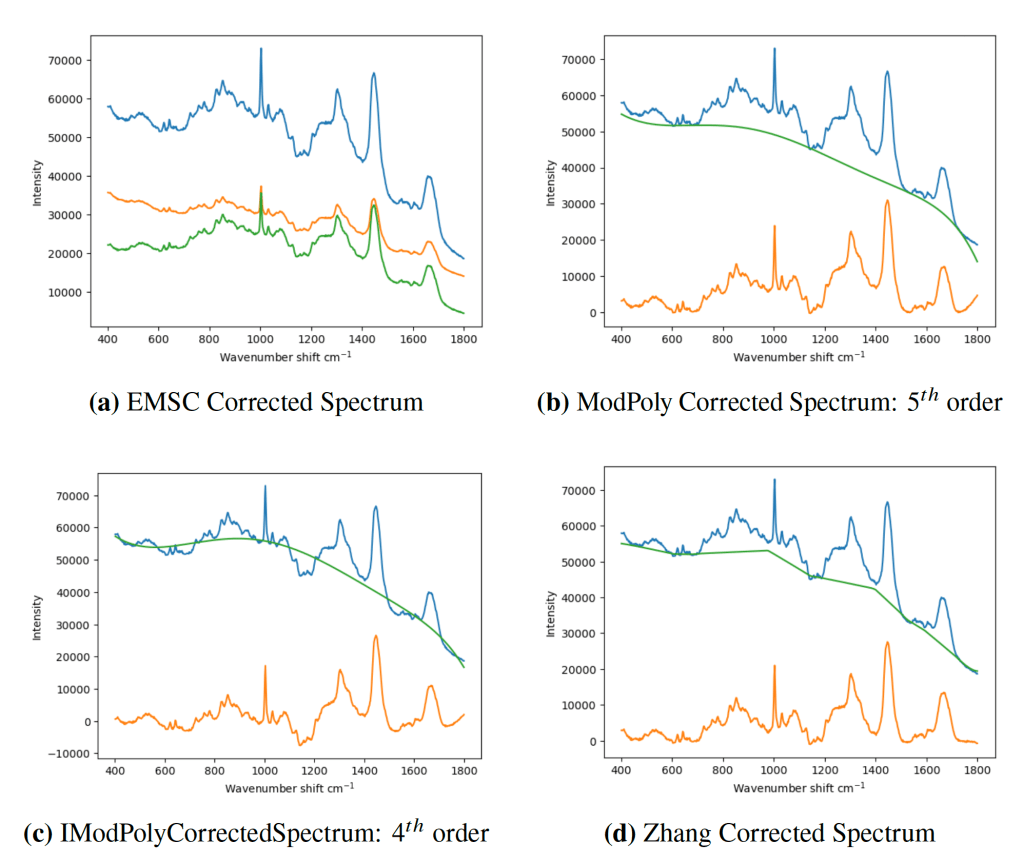
Figure 5.8 from my PhD. thesis. Blue is the original spectrum, green an estimated baseline and orange the ‘corrected’ spectrum. Which, if any, is the correct correction?
In the hope of finding a more objective method for choosing amongst this vast array of possibilities, I thought to look at downstream performance applying machine learning to a classification task. The literature was full of claims that baseline correction improved performance. At least then you could justify your choice as being the most diagnostically useful. So I tried a small range of different methods and their impact on performance.
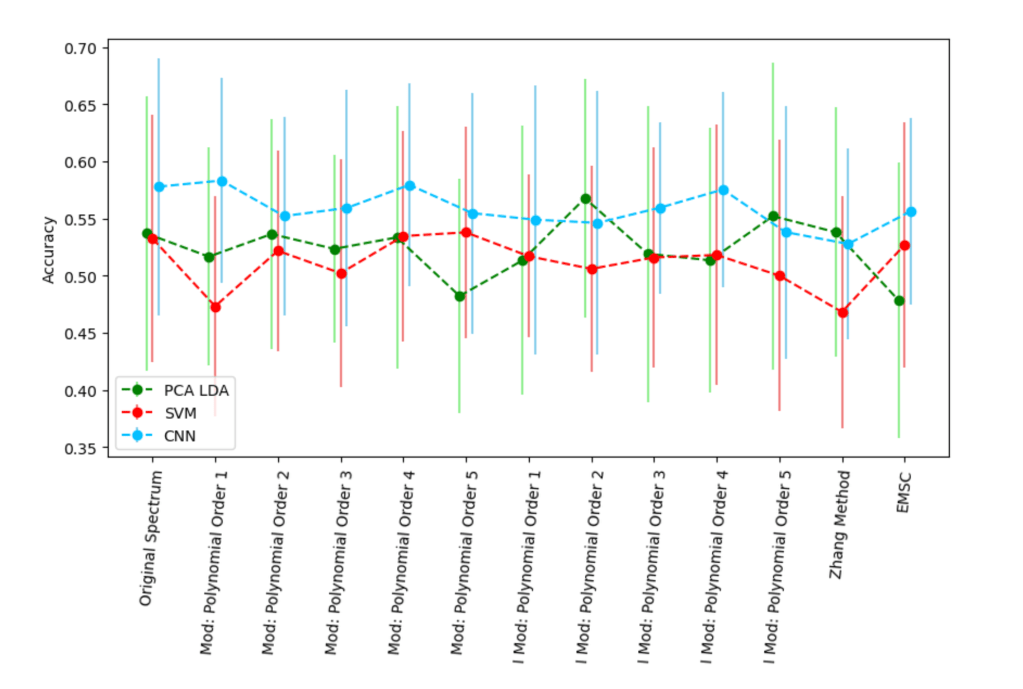
A small selection of baseline methods/hyperparameters and their resulting accuracy on a 5-class classification task on 3 machine learning models in otherwise identical training runs. Note those bars are 1 standard deviation – constructing confidence intervals from non-independent training folds is tricky.
The choice of baseline correction made no difference.
It is always possible to construct a training run to make any number you want go up. But you are just mining noise in the data. Most machine learning courses (I guess they’re all called AI courses now) will warn against the perils of over-fitting your data. However, this applies not just to the model parameters but also the model’s hyperparameters (e.g. learning rate) AND any pre-processing steps and their associated hyperparameters. Hence, you must treat hyperparameter selection, such as a the order of a polynomial baseline during correction, as you would model hyperparameter selection, using techniques such as nested cross-validation
Bocklitz and Popp, big names in biomedical spectroscopy, looking in far more detail found something similar. They investigated some 30000 possible pre-processing combinations (including baseline correction methods/hyperparameters) and found that very few slightly improved performance. This is huge space to search, and with no principled way of searching, you are far more likely to harm performance rather than increase it. Even though a poorly designed experiment will show an improvement, all that has really been achieved is that the generalisation gap has been increased. This is the modern incarnation of p-hacking. Make number go up =/= good.
This is why I do not baseline correct my spectroscopy data. Granted I’m only interested in disease classification, and there are other tasks which may benefit from careful baseline correction. But I believe the principle of parsimony also includes altering your data as little as possible. In the context of the deep learning paradigm there’s even less reason to baseline correct – give your model enough data (OK, easier said than done), then it will learn what function(s) needs to be applied to the data to minimise loss, which may or may not include a function to remove a baseline.
This was a bit of a rant – I still get reviewers asking why I didn’t baseline correct. I think the default question should be why did you baseline correct, and why in this way.



Leave a comment